|
Project data are stored in Mendeley data repository. Access can be gained below:
Spring 2018 Data Summer 2018 Data |
This research project began in March 2018 and has been through several iterations. Over the past four years, each cohort of students has chosen to pursue the same general questions regarding the abundance of antibiotic (ampicillin) resistance genes in soils from different land-use locations (crop-lands used for agriculture and soils from areas adjacent to rivers and streams; local parks, main campus areas). They have hypothesized that there should be more antibiotic resistant bacteria and therefore more resistance genes in soils around flowing water sources. They think this would be a consequence of watershed drainage systems having input of runoff from surrounding areas, especially in low-lying creek and river bottoms.
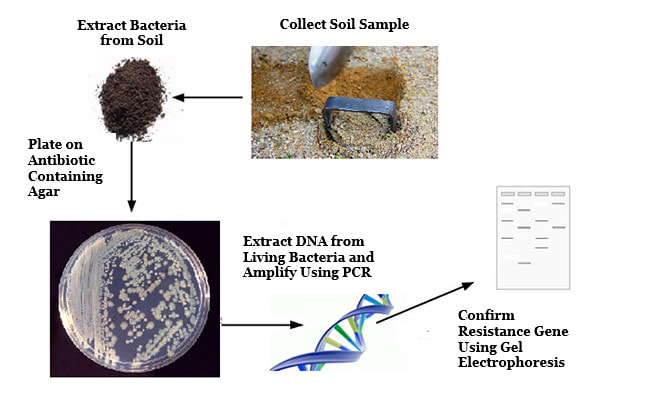
Each student is randomly assigned a set of coordinates to serve as their collection point. Utilizing Google maps, students locate their collection point and collect a soil sample. We will be following the protocol for detection presented by Bell et al. 2016. After determining the presence of any ampicillin resistance genes, we may further test for susceptibility to other antibiotics (penicillin, erythromycin, streptomycin, and tetracycline), using the basic Kirby-Bauer testing method for the first time during the project. |
Photo used under Creative Commons from NIAID